Updates
- 8/30/2012
-
Release of MATS 3.0.2.beta,
-
Fixed a bug related to default insert sizes and standard distribution.
|
|
|
- 8/24/2012
-
Release of MATS 3.0.1.beta,
-
A bug related to Ubuntu (tested on 12.04) is fixed.
|
|
|
- 8/9/2012
-
Release of MATS 3.0.0.beta, a major update with important features added to MATS:
-
Replicate data support: MATS now works with replicate RNA-Seq data from both paired and unpaired study design.
|
-
Improved filtering system: MATS now filters out AS events where large gene expression fold changes may confound the analysis.
|
-
MacOS support: MATS now supports both Linux and MacOS (tested on MacOS 10.8.1).
|
|
|
- 5/25/2012
-
Release of MATS 2.1.0
-
MATS now works with both read sequence files (fastq) and mapped reads files (bam). Using bam files
adds flexibility in mapping because MATS will skip the read mapping step.
|
-
A bug related to an empty AS event file is fixed.
|
|
|
- 5/15/2012
-
Release of MATS 2.0.0, a major update with important features added to MATS:
-
Simplified running procedure: MATS now only requires the raw RNA-Seq data, a genome sequence file, and a gene/transcript annotation file in GTF format as the input.
|
-
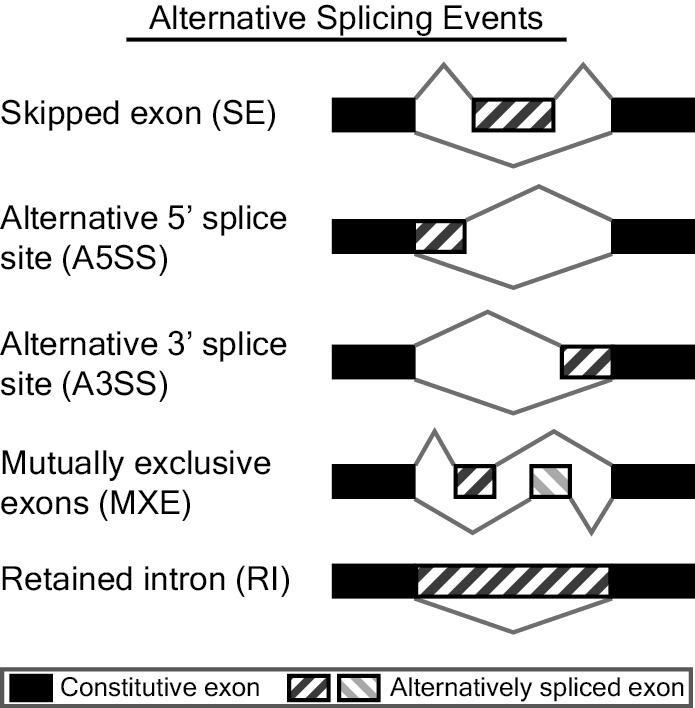
Ability to analyze different types of alternative splicing events: MATS now automatically detects and analyzes alternative splicing events corresponding to all major types of alternative splicing patterns.
|
-
Improved statistical power: MATS now works with both exon-exon junction reads and exon body reads which leads to improved statistical power.
|
|
|
- 3/5/2012
- Release of MATS 1.2.0, added a new method to calculate P-values by likelihood-ratio test, which is ~100x faster than the Bayesian method.
|
|
- 2/16/2012
- Release of MATS 1.1.0, adds Ensembl version of mouse annotation.
|
|
- 12/15/2011
- Release of MATS 1.0.0, the initial version of MATS was opened.
|
|
Contact
Correspondences regarding the MATS algorithm should be directed to Prof. Yi Xing (yxing at ucla.edu) and Shihao Shen (shihao-shen at uiowa.edu).
Correspondences regarding running of the MATS pipeline should be directed to Juw Won Park (jwpark2012 at ucla.edu).
Citation
Shen S., Park JW., Huang J., Dittmar KA., Lu ZX., Zhou Q., Carstens RP., Xing Y.
MATS: A Bayesian Framework for Flexible Detection of Differential Alternative Splicing from RNA-Seq Data.
Nucleic Acids Research, 2012;40(8):e61
doi: 10.1093/nar/gkr1291